Abstract
BACKGROUND AND PURPOSE: Although posttraumatic epilepsy is a common complication of traumatic brain injury, the relationship between these conditions is unclear and early posttraumatic epilepsy detection and prevention remain major unmet clinical challenges. This study aimed to identify imaging biomarkers that predict posttraumatic epilepsy among survivors of traumatic brain injury on the basis of an MR imaging data set.
MATERIALS AND METHODS: We performed tensor-based morphometry to analyze brain-shape changes associated with traumatic brain injury and to derive imaging features for statistical group comparison. Additionally, machine learning was used to identify structural anomalies associated with brain lesions. Automatically generated brain lesion maps were used to identify brain regions where lesion load may indicate an increased incidence of posttraumatic epilepsy. We used 138 non-posttraumatic epilepsy subjects for training the machine learning method. Validation of lesion delineation was performed on 15 subjects. Group analysis of the relationship between traumatic brain injury and posttraumatic epilepsy was performed on an independent set of 74 subjects (37 subjects with and 37 randomly selected subjects without epilepsy).
RESULTS: We observed significant F-statistics related to tensor-based morphometry analysis at voxels close to the pial surface, which may indicate group differences in the locations of edema, hematoma, or hemorrhage. The results of the F-test on lesion data showed significant differences between groups in both the left and right temporal lobes. We also saw significant differences in the right occipital lobe and cerebellum.
CONCLUSIONS: Statistical analysis suggests that lesions in the temporal lobes, cerebellum, and the right occipital lobe are associated with an increased posttraumatic epilepsy incidence.
ABBREVIATIONS:
- FDR
- false discovery rate
- ISLES
- Ischemic Stroke Lesion Segmentation
- ML
- machine learning
- PTE
- posttraumatic epilepsy
- ROC
- receiver operating characteristic
- TBI
- traumatic brain injury
- TBM
- tensor-based morphometry
- VAE
- Variational Autoencoder
The onset of posttraumatic epilepsy (PTE) after traumatic brain injury (TBI) is relatively common.1 Epidemiologic studies have found that PTE accounts for 10%–20% of all symptomatic epilepsies in the general population and ∼5% of all epilepsies.2 Significant risk factors for seizures occurring or continuing beyond 1 week after TBI include the occurrence of seizures within the first week, acute intracerebral (especially subdural) hematoma, brain contusion, greater injury severity, and age older than 65 years at the time of injury.3 As many as 86% of patients with 1 seizure early after TBI experience a second one within the next 2 years.4
Despite the reported relationship between TBI and PTE, identifying biomarkers of epileptogenesis after TBI is still a fundamental challenge. Preliminary studies in adult male Sprague-Dawley rats indicated the potential involvement of the perilesional cortex, hippocampus, and thalamus in PTE and demonstrated the potential of leveraging MR imaging analysis to find PTE biomarkers.4,5 Previous MR imaging studies have shown correlations between PTE incidence and the presence of lesions in T2-weighted scans, injury severity, and injury type.6⇓⇓-9 Studies of PTE reported correlations between PTE and the existence of frontal, parietal, and temporal lesions.10-12 Nevertheless, the association between PTE and lesion size or location remains poorly understood. Additionally, the heterogeneous nature of TBI injury types, pathology, and lesions presents additional challenges to biomarker discovery. Because the locations, spatial extent, and content of lesions vary considerably among patients with-versus-without PTE, there is no complete spatial overlap of injury profiles across the 2 groups. This heterogeneity needs to be accounted for in statistical analyses due to its potentially confounding effect. The prediction of posttraumatic seizure onset and frequency based on neurologic and radiologic examinations has been only moderately successful, and more research is needed to understand the relationship between TBI and PTE.10,13,14 Thus, the identification of imaging biomarkers can help in developing better PTE prediction strategies.
This study uses multimodal MR imaging from subjects with TBI to identify location- and contrast-related biomarkers for those who will develop PTE. We performed 2 analyses aimed at characterizing changes in brain structure using 2 distinct strategies:
Morphometric analysis: We performed a population analysis of morphometric changes in the brain associated with TBI. In contrast to the lesion analysis described below, this analysis focused on identifying changes in brain shape rather than alterations in tissue composition.
Lesion analysis: We used a machine learning (ML) method for identifying abnormal contrasts in multimodal MR images, which are indicative of lesion and tissue abnormalities such as edema, hematoma, and hemorrhage.
The morphometric and lesion analyses provide distinct-but-complementary information about the structure of the brain. Morphometric analysis focuses on the differences in brain shape, while the lesion analysis focuses on differences in tissue characteristics. TBI can cause both types of structural changes in the brain: changes in brain shape as a result of edema or direct injury as well as changes in brain tissue characteristics due to lesions. Therefore, it is important to analyze both of these aspects of structural brain change as a result of TBI.
MATERIALS AND METHODS
Data
We used 3 data sets in this study: 1) the Maryland TBI MagNeTs data set,15 2) the TRACK-TBI Pilot,16 and 3) the Ischemic Stroke Lesion Segmentation (ISLES) data set with manually delineated lesions.17 These data sets were used as follows: The neural network was trained with 97 subjects using Track-TBI Pilot and 41 subjects using MagNeTs. For validation of the neural network, we used 15 subjects from the ISLES data. Statistical analysis using morphometry and lesions (group differences) was performed using the MagNeTs data set, with a different subset of subjects from those used for training the neural network. We used 37 subjects with TBI who later developed epilepsy (26 males/11 females, 16–65 years of age) and 37 randomly selected subjects who did not (27 men/10 women, 18–70 years of age). In all TBI data sets, MR imaging was collected within 10 days of injury. More detailed information about these 3 data sets is provided in the Online Supplemental Data.
Preprocessing
Preprocessing of all 3 data sets was performed using the BrainSuite software (https://brainsuite.org). The 3 modalities (T1, T2, FLAIR) were coregistered to each other by registering T2 and FLAIR to T1. The T1 images were also each coregistered to the Montreal Neurological Institute atlas using a rigid (translation, scaling, and rotation) transformation. As a result, we transformed images in all 3 techniques to the Montreal Neurological Institute atlas space at a 1-mm3 resolution. Brain extraction was performed by stripping away the skull, scalp, and any nonbrain tissue from the image using BrainSuite. This was followed by tissue classification and generation of the inner and pial cortex surfaces.
Lesion identification using a neural network was performed on 2D axial images. Following registration to the Montreal Neurological Institute atlas and removal of nonbrain tissue, all 2D axial images (T1, T2, FLAIR) were reshaped to 128 × 128 pixels and histogram-equalized to a lesion-free subject. These data were then used for training and testing the lesion-detection network.
For tensor-based morphometry (TBM) and volumetric lesion analysis, we performed a further deformable registration of all subjects to a common atlas. The extracted cortical surface representations and brain image volumes for each subject were jointly registered to the BCI-DNI Brain Atlas (http://brainsuite.org/svreg_atlas_description). This coregistration establishes a one-to-one correspondence between individual subjects’ T1 MRIs and the atlas. The deformation map that transforms between the subject and the atlas encodes any morphometric differences between subject and atlas.
To achieve this registration, BrainSuite first performs a series of processing steps that involve correction for image contrasts as well as other scan and structural anomalies. These involve bias field correction that corrects for tissue-contrast changes due to field inhomogeneities during MR imaging. Other stages in the processing sequence include anisotropic filtering, topology correction, and dewisping modules to remove small errors due to noise and limited scan resolution. The Surface-Volume Registration module18 in BrainSuite then performs coregistration of individual subjects to an atlas. The volumetric coregistration in Surface-Volume Registration is constrained by cortical surface matching, to ensure 1–1 alignment of the subject and atlas cortices. The incorporation of surface and volume matching constraints makes the coregistration robust to the presence of lesions and missing or abnormal brain tissue as a result of injury. As a result, the correspondence between subject and atlas at the lesion is interpolated on the basis of the surrounding tissue, and anatomic correspondence away from the anomalies is minimally affected. Additionally, the results of the BrainSuite processing sequence were manually inspected to ensure accurate coregistration and exclude failed processing.
Tensor-Based Morphometry
To perform a morphometric analysis that compares the brain shapes of patients with PTE with those of participants without PTE, we used TBM.19 TBM is an established neuroimaging method that identifies regional differences in brain structure in groups or individuals relative to a control group using the determinant of the Jacobian matrix computed from the deformation field; the latter defines a nonlinear mapping that warps the brain into a common (atlas) space.20 Regions of the brain that differ most from the reference atlas brain will be characterized by significantly smaller (eg, atrophy/tissue loss) or larger (eg, enlarged ventricles) Jacobian determinants relative to controls. For this group study, we used 37 subjects with TBI who later developed PTE and 37 who did not from the MagNeTs data set.
We used the TBM pipeline in BrainSuite to map structural brain changes resulting from TBI to identify regions that are more strongly associated with the onset of PTE.19,21 The Jacobians are computed from the deformation fields associated with the cortically constrained volumetric subject-to-atlas registration described above. We applied 3-mm SD (7-mm full width at half maximum) isotropic smoothing to the Jacobian determinant maps to account for residual misregistration and to increase statistical power.
We analyzed the Jacobian determinants at each voxel using an F-test to determine whether there were group differences in the variances of this measure. The null hypothesis for the test is that the variances of Jacobian determinants in the PTE and non-PTE groups are equal. Our reason for applying the F-test is as follows: Because trauma affects different areas in the brain in different subjects across groups, it is unlikely that consistent localized differences between the 2 groups would be observed. Accordingly, we found that a t test of differences in the group means did not show significance. Because there may be >1 region in which lesions lead to a higher probability of developing PTE, we hypothesized that only a subset of subjects with PTE would have TBI-related differences from the non-PTE group in any particular area, leading to a larger variance across the PTE group in these areas relative to the non-PTE group. Thus, we performed the F-test, which allows us to observe larger variances in localized shape differences in the PTE relative to non-PTE group in regions at higher risk for developing PTE foci. The resulting P values were corrected for multiple comparisons using the Benjamini-Hochberg false discovery rate (FDR) procedure.22
Lesion-Based Analysis
To complement the TBM analysis, which captures morphometric brain changes, we also performed a lesion-based analysis to analyze changes in the underlying tissue microstructure, edema, and other TBI-related factors revealed by MR imaging contrast changes. For lesion mapping, we used multimodal MR images (T1, T2, FLAIR) and ML to automatically identify and delineate abnormal tissues. Lesions can be identified by visual inspection after extensive training, but this time-consuming process makes ML an attractive alternative. Approaches based on supervised ML have already achieved noticeable success, reaching high accuracy for lesion detection.23,24 Many manual lesion delineations are required to train supervised machines. In contrast, unsupervised approaches do not require labeled data but can be less accurate. Results are presented below reporting on the accuracy of the unsupervised method used here for lesion detection.
A popular unsupervised ML approach to lesion identification leverages a form of deep learning neural network known as a Variational Autoencoder (VAE).24,25 The VAE is a directed probabilistic graphic model whose posteriors are approximated by a neural network. By training the VAE using nominally normal imaging data, the network learns to encode only images with normal findings. As a result, the associated image “decoder” can reconstruct these images. When presented with images containing lesions or anomalies, the VAE encodes and reconstructs the image as if it contained only normal structures, as illustrated in Fig 1. Lesions as well as other pathology that may include hematoma, edema, and hemorrhage can then be identified from the differences between original and VAE-decoded images.
The VAE network and an input/output sample pair from the ISLES data set. X denotes the input data; Z denotes its low-dimensional latent representation. The VAE consists of an encoder network that computes an approximate posterior qφ (Z|X), and a decoder network that computes pθ (X|Z). The VAE model takes T1, T2, and FLAIR images from individual subjects (left), compresses them to generate a latent representation (Z), and regenerates 3 images (right). The VAE is trained on a data set that contains few lesions. After training, when presented with a newly lesioned brain, the reconstruction effectively removes the lesion from the image, resulting in a normal (lesion-free) version of the brain.
For the architecture of the VAE, we used the convolutional neural network proposed in Larsen et al 2015,26 which consists of 3 consecutive blocks of convolutional layers, a batch normalization layer, a rectified linear unit activation function, and 2 fully connected layers in the bottleneck for the encoder. For the decoder, we used a fully connected layer and 3 consecutive blocks of deconvolutional layers, a batch normalization layer, and a rectified linear unit with a final deconvolutional layer (Fig 1). The VAE detects lesions in 2D axial images of 128 × 128 pixels. The size of the input layer is 3 × 128 × 128, accommodating T1, T2, and FLAIR data. A more detailed description of this method and architecture is available in Akrami et al.24 Lesions are delineated on the basis of the VAE error between the input and reconstructed FLAIR images. Volumetric lesion maps are re-assembled from these 2D images. The resulting 3D VAE lesion maps are then warped to the BCI-DNI atlas space by applying the deformation field computed to map each subject to the atlas as described above. By representing all lesion maps in a common atlas space, we are then able to perform the statistical analysis described below.
We used a combination of 97 subjects from the Maryland TBI MagNeTs data set and 41 from the TRACK-TBI Pilot data to train the VAE. These data sets are not lesion-free, but a VAE can handle occasional lesions in the training set because it has some degree of robustness to outliers. To ensure this, we compared its performance with our recently described Robust-VAE24,27 and confirmed that there was no significant difference between their results. Despite this robustness, the number of anomalies in the training data should be minimized, and because we expected the lesion load to be somewhat lower in the non-PTE group, we used only subjects without PTE for training. Validation of lesion delineation using the trained VAE was performed using 15 subjects from the manually labeled ISLES data.
A group study of the relationship between lesion load and location and PTE onset was performed using the same 74 subjects (37 with and 37 without PTE) as in the TBM study. Using a method similar to the TBM analysis, we analyzed the VAE lesion maps using an F-test to determine whether there were statistically significant differences in the variances of lesion maps between the PTE and non-PTE TBI groups. As with the TBM analysis, we also confirmed that the t test did not show any significance due to the heterogeneity of lesion locations. Because lesion locations vary across subjects, some subjects in either group have healthy-appearing tissue at a given location in the brain, whereas some have lesions. However, if lesions in a brain region increase the chance of PTE, then in that region, we would expect to see greater heterogeneity across the PTE than the non-PTE group, leading to an increase in variance as reflected in the F-test on the lesion maps. The resulting P values were corrected for multiple comparisons using the Benjamini-Hochberg FDR procedure.22
We also performed a regional analysis by quantifying lesion volume from binarized lesion maps in each lobe using our USCLobes brain atlas (http://brainsuite.org/atlases/). This atlas consists of lobar delineations (left and right frontal, parietal, temporal, and occipital lobes, as well as the bilateral insula and cerebellum). To identify the lesions using a binary mask in each lobe, we applied a 1-class support vector machine28 to the VAE lesion maps at each voxel and across subjects to identify subjects with abnormally large VAE reconstruction errors at that voxel. The 1-class support vector machine is a commonly used unsupervised learning algorithm for outlier detection.28,29 We used the outliers marked by the 1-class support vector machine as lesion delineations and computed lesion volumes per lobe by counting the number of outlier voxels in each lobe for each subject.
Validation of VAE Lesion Detection
After training the VAE, we evaluated its performance using 15 subjects from the ISLES data set for which manually delineated lesions are also available. We calculated the pixel-wise absolute reconstruction error and applied median filtering to the resulting image to remove isolated pixels. Ground truth was defined using hand-traced delineation of lesions on FLAIR images.17 We then generated receiver operating characteristic (ROC) curves and computed the area under the curve on the basis of the concordance between pixels in the labeled lesions and those pixels in which the absolute error image exceeded a given threshold. The ROC curves were generated by varying the lesion-threshold intensity in the error image to control the true- and false-positive rates.
RESULTS
TBM-Based Analysis
The results of TBM analysis using F-tests applied to the Jacobian determinant maps are shown in Fig 2. As anticipated, in the case of the t-statistic map (not shown), TBM analysis results did not survive multiple-comparison corrections for the FDR using the Benjamini-Hochberg procedure.22 This result may be because of the heterogeneity of lesion locations and sizes across both groups. In contrast, the F-test is sensitive to significant differences in variance between the 2 groups and does show regions where the Jacobian determinant is significantly different, even after FDR correction (q = 0.05). The voxels close to the pial surface associated with significant differences may indicate group differences in the locations of edema, hematoma, or hemorrhage and may, therefore, be associated with an increased risk of PTE.
Three orthogonal views through the P values thresholded at P = .05 (FDR-corrected) obtained for the F-test for TBM analysis using Jacobian determinants. L indicates left; R, right.
Lesion-Detection Performance
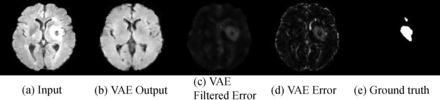
Performance of the VAE lesion-detection methods was quantified using ROC analysis on 15 subjects from the ISLES data set. Due to the infrequent occurrence of lesions in the training data, the VAE was able to prevent reconstruction of lesions so that they appeared in the error map. We illustrate VAE performance for cases in which lesions are present in Fig 3. Note that the reconstructed images in B are “de-lesioned” approximations of the input images in A. Normal tissue is reconstructed, whereas anomalies and lesions are not. The error maps in C are indicative of anomalies in the brain. The error maps after median filtering in D show reasonable correspondence with the ground truth E. The area under the curve for the lesion-detection ROC study on the ISLES data was 0.81 (SD, 0.003) (the confidence interval was achieved using 100 bootstrap iterations). We also calculated the Dice coefficient, which quantifies the intersection between 2 sets, in our case the ground truth and VAE-determined lesion volumes. To define lesion volume, we thresholded the VAE error at a false detection rate per voxel of 0.01. The average Dice coefficient across the test set was 0.47 (SD, 0.29).
Reconstruction results obtained by applying the VAE to the ISLES data set. A, Sample slices from input images. B, Slices reconstructed from the VAE. C, Difference between input and reconstructed images. D, Error maps after applying median filtering to reduce the occurrence of spurious voxels. E, Manually delineated lesion masks used as ground truth to evaluate VAE performance.
Lesion-Based Analysis
The results of the F-test showed significant differences between groups in both the left and right temporal lobes. We also saw significant differences in the right occipital lobe and the cerebellum (Fig 4). The results of the lobar analysis (Table) were consistent with voxelwise analysis, showing an increased variance in the PTE population relative to subjects without PTE in the left and right temporal lobes, right occipital lobe, and cerebellum.
Orthogonal views through the P values thresholded at P = .05 (FDR-corrected) obtained for the F-test comparing lesion maps for the PTE and non-PTE groups. L indicates left; R, right.
Lobe-wise lesion volumes as measured using a 1-class SVM to generate binary lesion maps
DISCUSSION
Our results are consistent with earlier TBI studies that showed a relationship between lesion location and the probability of PTE onset. In particular, the F-test in our lesion study indicates a correlation between PTE presence and the frequency of lesion occurrence in temporal lobes, consistent with previous studies.10⇓-12,30,31 Most interesting, the TBM F-test shows areas of significant differences between groups that are, in large part, clustered on or just below the pial surface as well as in the cerebellum. Furthermore, these TBM results appear largely complementary to the lesion analysis, indicating effects near the pial surface in contrast to the larger scale and deeper lesion-related findings. While the near-surface clusters could be false-positives and need further investigation, this result may indicate the increased occurrence of edema or hematoma in patients with acute TBI, which is known to alter the cortex shape32 and which may be associated with an increased chance for developing PTE.
TBM and its extensions20,33 have been used for whole-brain analysis of structural abnormalities in patients with temporal lobe epilepsy. Significant volume reductions were found in brain regions including the hippocampus, cingulate gyrus, precentral gyrus, right temporal lobe, and cerebellum.33 Cross-sectional studies of children with chronic localization-related epilepsy using traditional volumetric and voxel-based morphometry have revealed abnormalities in the cerebellum, frontal and temporal lobes, hippocampus, amygdala, and thalamus.34⇓⇓-37 The etiology of these cohorts involves mesial temporal sclerosis,34 and it may even be cryptogenic.35 Pediatric and adult populations present slightly different patterns in gray matter atrophy; however, the involved regions are largely common.36 One exception is epileptic encephalopathies in infants and children.38
While in mesial temporal lobe epilepsies, hippocampal sclerosis is the most common pathologic finding,39 neuronal damage is often not restricted to the hippocampus. MR imaging morphometric studies have demonstrated extrahippocampal and extratemporal atrophy in adults with mesial temporal lobe epilepsies.40⇓⇓-43 These studies also emphasized the role of temporal lobe damage in epilepsy, which provides further evidence for the role of temporal lobe lesions in PTE. The findings from our study further support this evidence for the involvement of the temporal lobe. Furthermore, studies like ours may assist or complement efforts to study posttraumatic metabolic crises44 or to localize posttraumatic epileptic foci for surgical resection via electroencephalography.45 The novel use of a VAE here to automatically delineate lesions may prove useful for future studies over the large data sets or collections of data sets like Federal Interagency Traumatic Brain Injury Research Informatics Systems (https://fitbir.nih.gov/), in which manual segmentation is very time-consuming and/or subject to large interrater variability.
One limitation of this study is that individualized clinical data related to the injury mechanism and severity of the injury, known risk factors for PTE,46 are not available in the public data sets available for research studies. Similarly, while the range of the Glasgow Coma Scale score, a predictor of PTE, is available, individualized Glasgow Coma Scale scores are not. Another limitation is that while recurrent seizures were noted in the questionnaire for this study, additional information about the seizure frequency and types is unavailable. Also, having access to a delineated set of TBI-related lesions rather than the ISLES data for subjects with stroke used for validation here would help better optimize the TBI lesion-detection neural network. Despite these limitations, the statistical analysis shows the role of the temporal lobe in PTE and demonstrates the utility of imaging-based early markers of PTE.
CONCLUSIONS
In this study, we investigated the relation of MRI structural bio-markers to development of epilepsy in posttraumatic patients. Our results demonstrate that lesions in the temporal lobes, cerebellum, and the right occipital lobe are associated with an increased posttraumatic epilepsy incidence. Furthermore, our TBM results appear largely complementary to the lesion analysis, indicating differences in brain morphometry near the pial surface, possibly associated with edema, hematoma, or hemorrhage, to be associated with increased risk for PTE.
Footnotes
This work is supported by Department of Defense grant W81XWH-18-1-061 and by National Institutes of Health grants R01 NS074980, and R01 EB026299. A.I. is supported by the National Institutes of Health grant R01 NS 100973, Department of Defense contract W81XWH-18-1-0413, and a Hanson-Thorell Family Research scholarship. Data used in the preparation of this article reside in the Department of Defense and National Institutes of Health–supported Federal Interagency Traumatic Brain Injury Research Informatics Systems (FITBIR) in FITBIR-STUDY 0000314 and FITBIRSTUDY 0000246.
This article reflects the views of the authors and does not reflect the opinions or views of the Department of Defense or the National Institutes of Health.
Disclosure forms provided by the authors are available with the full text and PDF of this article at www.ajnr.org.
Indicates open access to non-subscribers at www.ajnr.org
References
- Received July 23, 2021.
- Accepted after revision January 1, 2022.
- © 2022 by American Journal of Neuroradiology